Deep Learning for Pediatric Crohn's Disease
A deep learning approach for predicting pediatric Crohn's disease from histopathological whole slide images, with investigation of inter-site generalizability and site-specific artifact mitigation.
Deep Learning for Predicting Pediatric Crohn's Disease Using Histopathological Imaging
Sharma A.H., Lawlor B.W., Wang J.Y., Sharma Y., Sengupta S., Fernandes P., Zulqarnain F., May E., Syed S., Brown D.E.
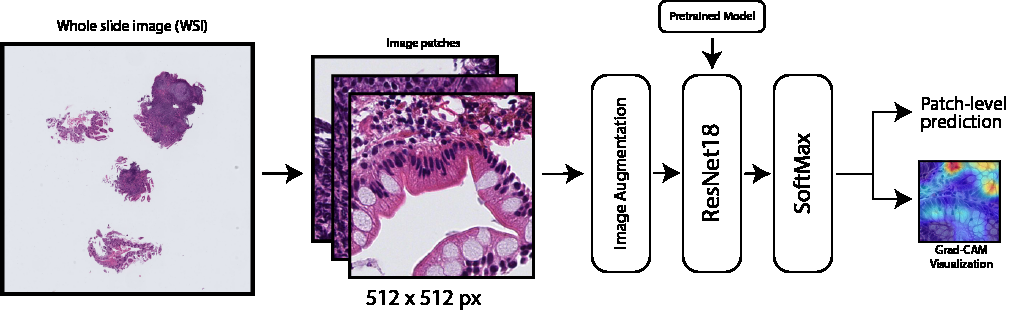
Architecture: WSI → 512x512 patches → Image Augmentation → ResNet18 → Patch-level prediction with Grad-CAM visualization
Overview
The current gold standard for Crohn's disease diagnosis involves examination of biopsied tissue by a trained physician, but endoscopic images and histological features are only evident when the appropriate biopsy site is chosen and the image is of high quality. This study developed patch-level image classification models using a ResNet18 CNN for prediction of Crohn's disease from H&E-stained whole slide images sourced from two hospital systems. While intra-site models achieved up to 93.9% accuracy, the discovery that a separate model could predict hospital source with 99% accuracy revealed significant site-specific artifacts — prompting investigation into stain normalization, image cropping, and other transformations to improve cross-site generalizability.
Abstract
Background
Crohn's disease is a chronic inflammatory bowel disease affecting the gastrointestinal tract, with pediatric patients typically experiencing a more severe disease course. The current gold standard for diagnosis involves manual examination of H&E-stained biopsies by trained pathologists, a process that takes approximately 30 minutes per patient and depends on biopsy site selection and image quality.
Objective
To develop a deep learning model for patch-level classification of Crohn's disease from whole slide histopathological images, and to investigate the generalizability of such models across different hospital systems where biopsy preparation, staining, and digitization protocols may differ.
Methods
We developed ResNet18-based binary classifiers trained on 512x512 pixel patches extracted from H&E-stained whole slide images from two hospitals: INOVA (n=91 patients) and Cincinnati Children's Hospital Medical Center (n=191 patients). Models were trained with weighted cross-entropy loss, color jittering augmentation, and ImageNet normalization. Grad-CAM visualizations were used for model interpretability.
Results
Intra-site models achieved 84.6% (INOVA) and 93.9% (CCHMC) validation accuracy. However, cross-site performance was poor — a separate model predicted hospital source with 99% accuracy, revealing site-specific artifacts. Stain normalization and random resized cropping reduced site-prediction accuracy to 74%, demonstrating partial mitigation of confounding features.
Significance
This study highlights a critical challenge for clinical deployment of histopathological deep learning models: site-specific artifacts from staining, tissue preparation, and digitization create confounding signals that prevent cross-site generalization. We identify specific sources of inter-site variation and propose transformations to reduce their impact, establishing a framework for developing more generalizable diagnostic models.
Challenges
- 1Whole slide images are extremely high-resolution (up to 110,000 x 110,000 px), prohibiting direct CNN application
- 2Site-specific artifacts from staining protocols, digitization, and tissue preparation create confounding features
- 3Imbalanced class distributions across hospital sites (CCHMC: 181 CD vs 10 control)
- 4Models must generalize across hospital systems to be clinically useful
Methodology
The pipeline transforms high-resolution whole slide images into patch-level predictions through a series of preprocessing, augmentation, and classification stages, with Grad-CAM-based interpretability analysis.
Data Collection
H&E-stained whole slide images of ileal biopsies from pediatric patients were sourced from INOVA (64 CD, 27 control) and Cincinnati Children's Hospital Medical Center (181 CD, 10 control). WSIs ranged from 20,000x20,000 to 110,000x110,000 pixels. Data was split 75:25 for training/validation with no patient overlap, and cross-site data served as test sets.
Image Patching
A sliding window method generated 512x512 pixel patches from each WSI with no overlap. A 50% white-space threshold filtered out patches with insufficient tissue. Each patch inherited the WSI-level label (CD or control), producing tens of thousands of training patches per site.
Image Augmentation
To address small sample sizes and reduce site-specific bias, color jittering was applied through random changes to brightness, contrast, saturation, and hue as a form of stain normalization. Random horizontal flips were applied, and RGB channels were normalized to ImageNet mean and standard deviation.
CNN Architecture & Training
ImageNet-pretrained ResNet18 served as the backbone for binary classification. Models were trained for 20 epochs using weighted cross-entropy loss with stochastic gradient descent (momentum 0.9), initial learning rate 0.001 decaying by 0.1 every 7 epochs.
Grad-CAM Interpretability
Gradient-weighted Class Activation Mapping was applied to the final convolutional layer to produce localization maps highlighting regions most important to each classification decision. Medical professionals reviewed Grad-CAM outputs to verify that models attended to clinically relevant histological features.
Inter-Site Investigation
A site-prediction model was trained to classify patches by hospital source. Stain normalization using StainTools and random resized cropping (128x128 from 512x512) were applied to systematically reduce site-distinguishing features. RGB channel distributions were analyzed to characterize staining differences between sites.
Approach
- 1Applied sliding window patching (512x512 px) with 50% white-space threshold filtering
- 2Fine-tuned ImageNet-pretrained ResNet18 with weighted cross-entropy loss and color jittering augmentation
- 3Used Grad-CAM visualizations for model interpretability, verified by medical professionals
- 4Investigated stain normalization (StainTools) and random resized cropping to reduce site-specific signatures
Results & Demos
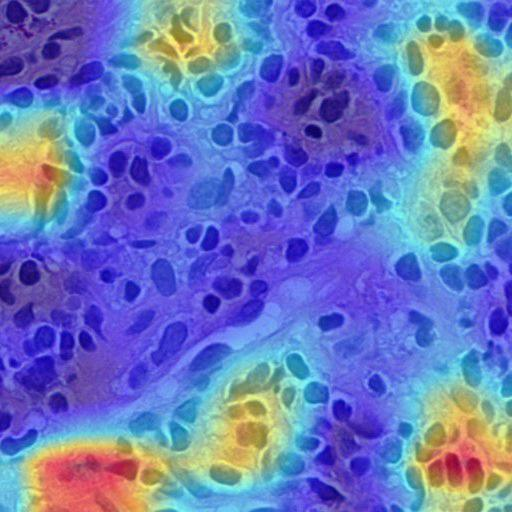
Grad-CAM: INOVA model predicted as control — highlighting lymphocytes in lamina propria
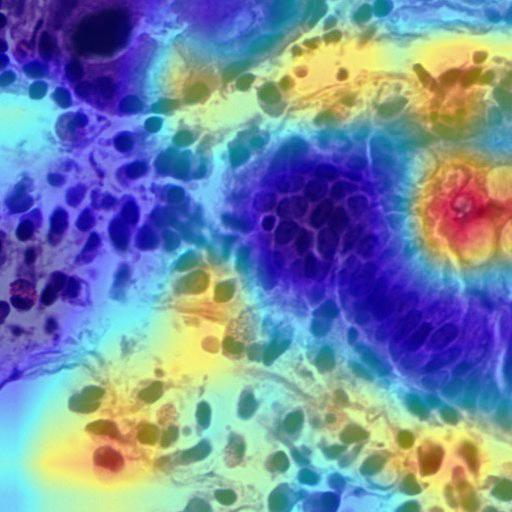
Grad-CAM: INOVA model predicted as Crohn's disease — highlighting goblet cells and white space
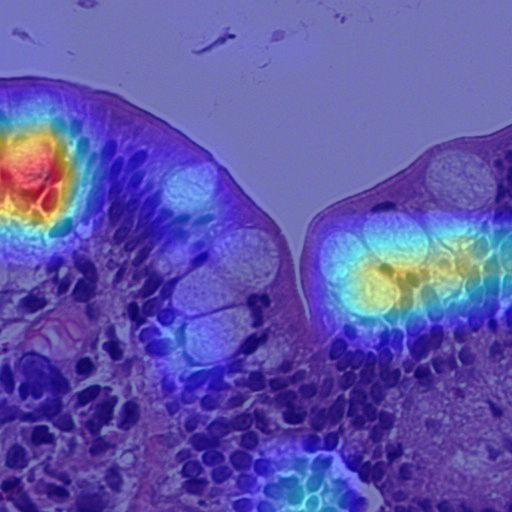
Grad-CAM: CCHMC model predicted as control — highlighting lymphocytes and occasional goblet cells
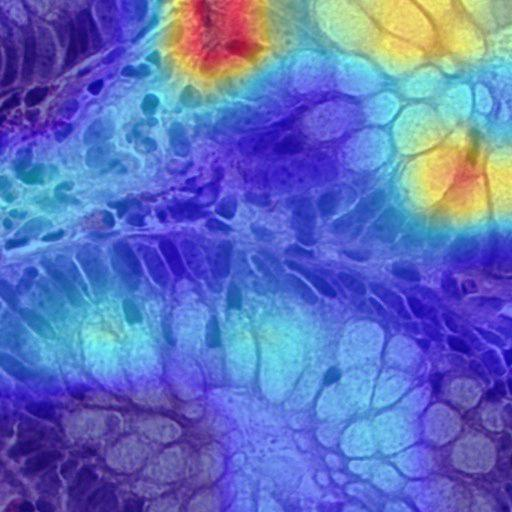
Grad-CAM: CCHMC model predicted as Crohn's disease — highlighting lymphocytes in lamina propria
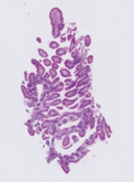
INOVA whole slide image showing increased white space relative to CCHMC
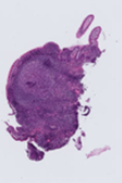
CCHMC whole slide image with denser tissue coverage
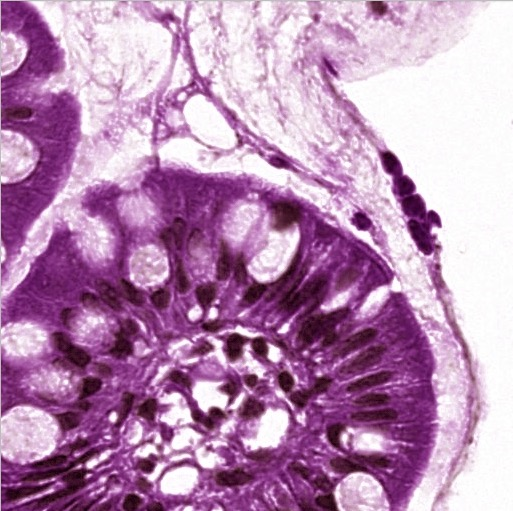
Biopsy before stain normalization — visible color variation between sites
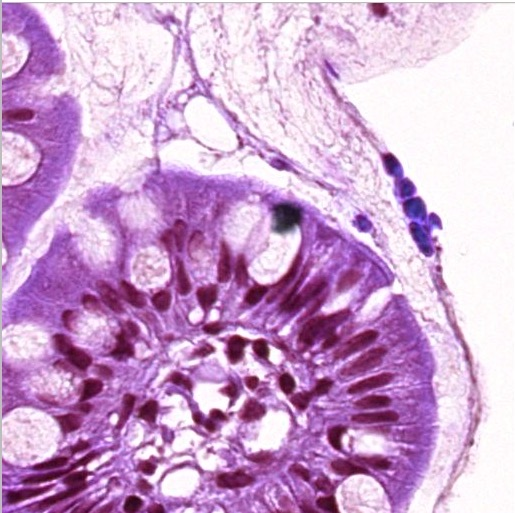
Biopsy after stain normalization using StainTools
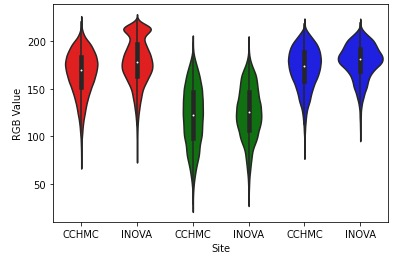
RGB channel distributions: red and blue channels differ between INOVA and CCHMC, reflecting H&E staining variation
Findings
Results demonstrate strong intra-site classification performance but reveal significant challenges for cross-site generalization due to detectable site-specific artifacts.
Intra-Site CD Prediction
Models trained and validated on same-site data achieved strong performance. Grad-CAM analysis confirmed that models focused on clinically relevant structures: lymphocytes in the lamina propria for control predictions, and goblet cells for CD predictions. The CCHMC model suffered from low precision due to class imbalance (181 CD vs 10 control).
| Site | Accuracy | F1-Score | Precision | Recall |
|---|---|---|---|---|
| INOVA | 84.6% | 0.843 | 0.767 | 0.937 |
| CCHMC | 93.9% | 0.506 | 0.367 | 0.812 |
Cross-Site Generalization Failure
When models were tested on data from the other hospital, performance dropped dramatically. The INOVA model achieved only 6.56% accuracy on CCHMC data, predicting nearly all patches as CD. The CCHMC model achieved 40.5% on INOVA data with similarly poor recall. This disparity motivated the investigation of site-specific features.
Site-Specific Artifact Detection
A model trained to predict hospital source from CD-only patches achieved 99% accuracy with no transformations applied. This confirmed the presence of strong site-specific signatures in the histological images that machine learning models readily exploit, even when such features are invisible to or routinely ignored by trained pathologists.
| Transformation | Site Prediction Accuracy |
|---|---|
| None (baseline) | 99% |
| Color jittering | 92% |
| Stain normalization (StainTools) | 94% |
| Color jitter + random resized crop | 74% |
RGB Channel Analysis
Comparison of RGB channel distributions revealed differences in the red and blue channels between sites, consistent with H&E staining variation. INOVA images showed a bimodal red channel distribution while CCHMC followed a unimodal distribution. The blue channel from INOVA had a smaller standard deviation compared to CCHMC, suggesting systematic differences in staining protocols.
Key Outcomes
- Achieved 84.6% (INOVA) and 93.9% (CCHMC) intra-site validation accuracy for CD prediction
- Identified site-specific artifacts detectable with 99% accuracy by machine learning models
- Reduced site-prediction accuracy from 99% to 74% through combined color normalization and image cropping
- Grad-CAM analysis confirmed model attention on clinically relevant structures (lymphocytes, goblet cells)
Discussion
The results reveal both the promise and critical limitations of deep learning for histopathological diagnosis, with important implications for clinical deployment.
The Generalizability Challenge
For WSI classification models to become useful in clinical settings, they must generalize across hospital systems. Our study revealed that site-specific artifacts — from staining protocols, tissue preparation, and digitization — create confounding signals that current CNN architectures readily learn. Pathologists naturally ignore these artifacts and focus on diagnostically relevant features, but deep learning models lack this domain knowledge without explicit intervention.
Artifact Mitigation Strategies
Two complementary approaches exist for addressing site-specific artifacts: removal through standardization of biopsy, slicing, staining, and imaging protocols; and model-awareness through training on diverse multi-site data. Our experiments showed that color normalization alone is insufficient (reducing site prediction only to 92-94%), while combining color normalization with geometric transformations (random resized cropping) more effectively reduced distinguishability to 74%.
Clinical Integration Path
The ultimate goal is to integrate machine learning into the pediatric CD clinical workflow to improve diagnostic accuracy and reduce physician labor. Achieving this requires models that generalize reliably across sites. Increased access to WSI datasets from diverse institutions, combined with synthetic data augmentation and domain adaptation techniques, would enable neural networks to learn about and overcome site-specific artifacts through training rather than preprocessing.
Read the Full Paper
Deep Learning for Predicting Pediatric Crohn's Disease Using Histopathological Imaging
Download Paper